Mitochondrial ribosomes
Mitochondria, which perform aerobic respiration to provide eukaryotic cells with ATP, evolved from free-living bacteria by endosymbiosis. Because of this evolutionary history, mitochondria still contain their own genome, as well as the molecular machines required to express the information encoded on it. Mitochondrial ribosomes (mitoribosomes) translate mitochondrial mRNAs into proteins. Most of the proteins that are synthesized by mitoribosomes - or in mammals, all of them - are membrane protein components of the complexes of the respiratory chain. In addition to their functional specialization, mitoribosomes have diverged markedly from their bacterial progenitors in terms of their structure and composition. Mitoribosomes are of great medical interest because antibiotics that interfere with their function can cause severe side effects, such as aminoglycoside-induced ototoxicity, and because defects in mitochondrial translation have been implicated in human hereditary diseases. Basil Greber's postdoctoral studies in the laboratory of Prof. Nenad Ban at ETH Zurich were focused on the determination of the complete structure of the 55S mammalian mitoribosome and its 39S and 28S mitoribosomal subunits.
The architecture of the mammalian 39S subunit
|
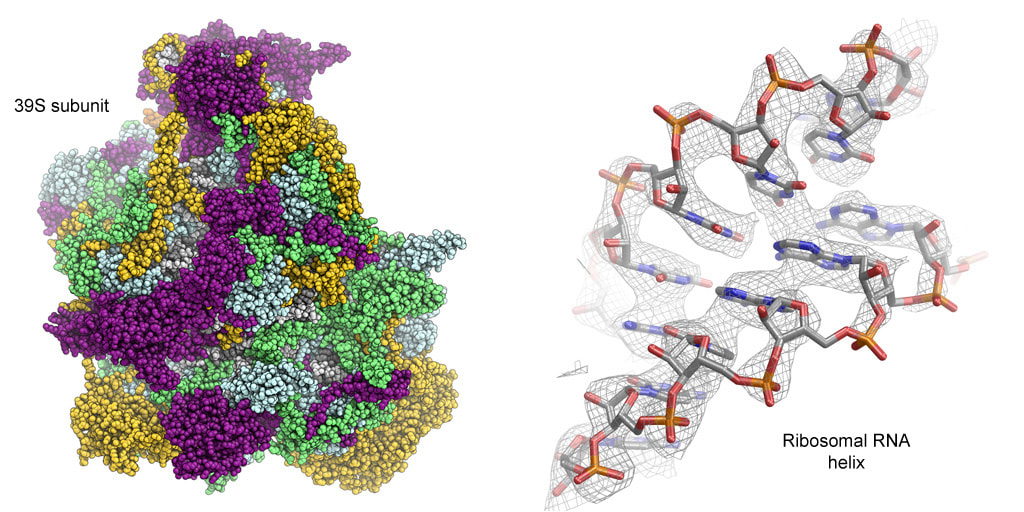
Cover page of Nature (left) featuring the mitoribosome, and 4.9 Å cryo-EM map of the mammalian 39S mitoribosomal subunit (right).
Greber BJ*, Boehringer D*, Leitner A, Bieri P, Voigts-Hoffman F, Erzberger JP, Leibundgut M, Aebersold R, Ban N (2014). Architecture of the large subunit of the mammalian mitochondrial ribosome. Nature 505 (7484): 515–519. Epub 2013 Dec 22. *equal contribution PubMed |
Using cryo-electron microscopy (cryo-EM) data collected using a direct electron detector camera and a FEI Titan Krios transmission electron microscope, we were able to calculate an cryo-EM density map of the 39S subunit at 4.9 Å resolution. Supported by chemical crosslinking-mass spectrometry (CX-MS) data obtained by Alexander Leitner in the group of Ruedi Aebersold at ETH Zurich, we were able to assign the locations of several mitochondrial-specific proteins. Our findings provided exciting insights into the membrane attachment of mitoribosomes as well as their molecular evolution.
The complete structure of the 39S subunit
Cryo-EM data collected using a second generation direct electron detector camera and the use of RELION for structure calculations enabled us to obtain a cryo-EM map of the 39S subunit in the context of the full 55S mitoribosome at 3.4 Å resolution. This reconstruction allowed the tracing of all proteins of the 39S subunit, with the exception of the highly flexible stalk regions, resulting in a fully refined and validated structure of the 39S subunit. Our structure revealed the presence of a tRNA molecule as an architectural component in the 39S central protuberance, structurally replacing the 5S rRNA found in a similar location in bacterial ribosomes. Additionally, our structure provided detailed insights into the conserved structure of the mitoribosomal peptidyl transferase active site bound with tRNA ligands as well as the architecture of the polypeptide exit tunnel.
|
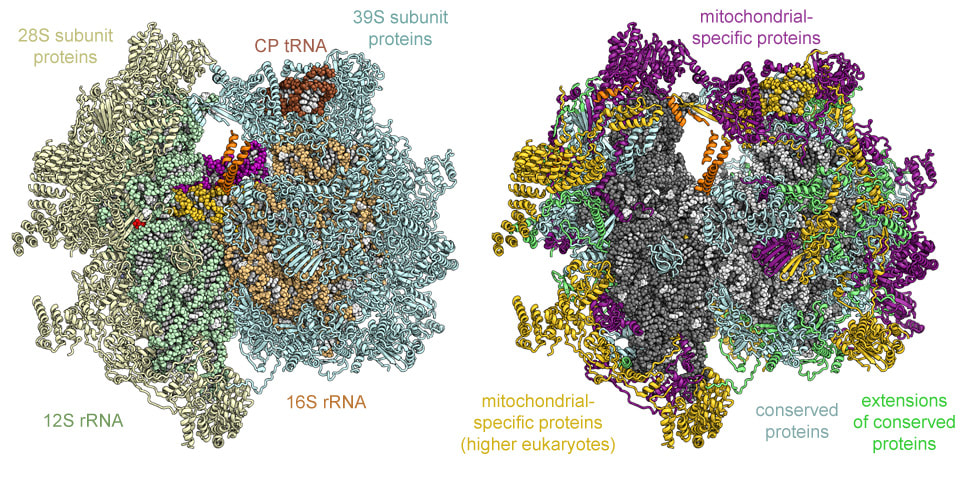
Left: Atomic coordinate model of the 39S subunit colored according to conservation. Mitochondrial-specific protein elements (green, purple, and gold) almost fully cover the solvent exposed side of the 39S subunit. Right: Example for the excellent quality of our cryo-EM density at 3.4 Å resolution (right).
Greber BJ*, Boehringer D*, Leibundgut M*, Bieri P, Leitner A, Schmitz N, Aebersold R, Ban N (2014). The complete structure of the large subunit of the mammalian mitochondrial ribosome. Nature 515 (7526): 283–286. Epub 2014 Oct 1. *equal contribution PubMed |
Structure of the mammalian 55S mitoribosome with bound mRNA and tRNA
|
Left: Computational rendering of the complete structure of the 55S mitoribosome including tRNAs (P-site purple, A-site gold) and mRNA (red). Right: Conservation of the protein components of the 55S mitoribosome.
Greber BJ*, Bieri P*, Leibundgut M*, Boehringer D, Leitner A, Aebersold R, Boehringer D, Ban N (2015). The complete structure of the 55S mammalian mitochondrial ribosome. Science 348 (6232): 303-308. Epub 2015 Apr 2. *equal contribution PubMed Greber BJ, Ban N (2016). Structure and Function of the Mitochondrial Ribosome. Annual Review of Biochemistry 85 (1). Epub 2016 Mar 24. PubMed |
Further computational sorting and analysis of the dataset that previously yielded the detailed structure of the 39S subunit resulted in a 3D reconstruction of the mitoribosomal 28S small subunit at 3.6 Å resolution and structures of the entire 55S mitoribosome, complete with mRNA and tRNA ligands, at 3.8-4.2 Å resolution. These cryo-EM reconstructions were of sufficient quality for manual model building, allowing us to assign the locations and folds of all proteins of the 28S subunit. The structure provides detailed insights into the highly divergent architecture of the 28S subunit, the structure of the mitoribosomal intersubunit bridges, the structure of the guanine-nucleotide binding protein mS29 at the head domain of the mitoribosomal small subunit, and the interactions of the 55S mitoribosome with its mRNA and tRNA ligands.
For additional information on the mammalian mitoribosome, including PyMol scripts for structure visualization and a complete guide to the nomenclature of mitoribosomal proteins, please visit the website of the laboratory of Prof. Nenad Ban at ETH Zurich.
For additional information on the mammalian mitoribosome, including PyMol scripts for structure visualization and a complete guide to the nomenclature of mitoribosomal proteins, please visit the website of the laboratory of Prof. Nenad Ban at ETH Zurich.